When considering a theory, we should first look at how successfully the theory is at predicting evidence. In my previous post, here, I discussed how Max Telford — an evolutionary biologist at University College London — relies on several foundational assumptions in his recent book, The Tree of Life: Solving Science’s Greatest Puzzle. These are assumptions that evolution critics, including myself, find unconvincing, illogical, or inadequately justified. However, in his narrative and to his credit, Telford also discusses several failed predictions from common descent. While he doesn’t call them “failed” predictions, he highlights them as important towards our modern understanding of the tree of life — and they show where ideas about common ancestry have led to bad predictions in the past. Let us consider:
- Prediction 1: Organisms pass through more primitive stages of evolution as they develop.
According to this evolutionary prediction, Ernst Haeckel famously — and notoriously — altered his drawings of animal embryos to better fit the expected pattern (p. 19). This same expectation drove researchers to search for a “lizard-like embryonic stage” in penguin development, a hypothesis that proved dramatically incorrect and is recounted on pp. 61–62 by Telford, including the near-fatal dangers encountered by the expedition members who pursued it. This prediction also led scientists to initially interpret the small brownish worm Xenoturbella as a “strange-looking mollusc.” A key supporting claim was that “Xenoturbella embryos, growing in the epidermis of their parent, pass through a stage that looks just like an oyster larva before metamorphosing into an adult” (p. 191).
Telford himself played a role in overturning this mistaken grouping. Yet, in offering a conciliatory explanation for why earlier researchers held this view, Telford writes: “This is certainly an unusual idea, but then a human baby grows right inside its mother’s body; and the human embryo passes through a stage with fish-like gill buds and a tail before changing into something tailless and very un-fish-like” (p. 191). Does this statement suggest that Telford continues to view the fish-like gill arches and tail in the human embryo as evolutionary holdovers from our ancestral past, rather than as features serving a functional purpose specific to that developmental stage? It certainly seems like he does hold this view — despite the fact that many mainstream scientists have criticized the old adage that “ontogeny recapitulates phylogeny.”
A Productive Prediction?
The question I invite the reader to consider is this: Has the prediction from evolutionary theory that embryos pass through ancestral stages proven productive? Has it genuinely advanced our understanding of embryonic development, or has it more often guided researchers down speculative paths that turned out to be incorrect?
- Prediction 2. Environmental pressures have shaped organisms through time leading to very different species being on the earth today as in the past. “The number of changes is closely correlated with how much time has passed” (p. 41).
Evolutionary theory predicts that over vast stretches of geological time (hundreds of millions of years), lineages are highly likely to undergo significant morphological, physiological, and genetic change due to natural selection, genetic drift, mutation, and environmental pressures. As a result, organisms that existed long ago are not expected to persist into the present day with little to no detectable modification. This is why it was such a surprise when a coelacanth, a “fossil” fish with a 400-million-year-old history, thought to be extinct for over 60 million years, was found alive off the coast of South Africa (p. 44)! Other well-documented examples of organisms showing remarkable stasis over hundreds of millions of years include:
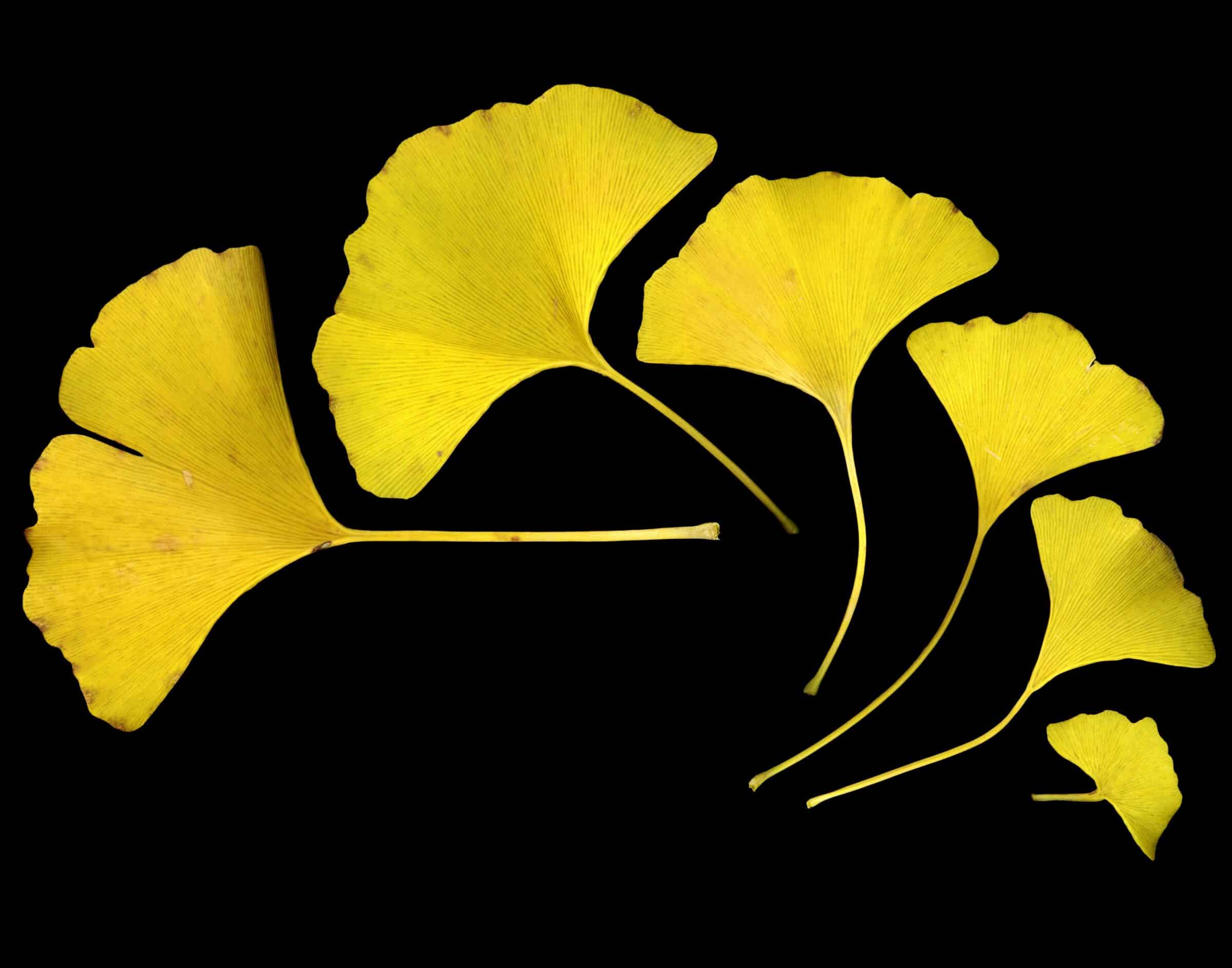
- Ginkgo trees — first appear in the fossil record during the Permian (~270 million years ago).
- Horseshoe crabs (Limulidae) — first appear in the Ordovician (~450 million years ago).
- Wollemi pine (Wollemia nobilis) — fossils date back to the Jurassic (~200 million years ago).
- Tuatara (Sphenodon punctatus) — lineage dates to the Triassic (~245 million years ago).
- Nautilus — chambered-shell morphology dates back to the Cambrian (~500 million years ago).
Difficulties trom the Molecular Clock
Another significant problem posed by these so-called “living fossils” relates to the molecular clock method of evolutionary biology.
The molecular clock assumes that genetic changes accumulate at a predictable, steady rate across lineages. As Telford explains: “If we measure how different two species are, as long as the rate of change is predictable, we should be able to calculate how distantly related (in years) they must be, that is how long ago they had a common ancestor” (p. 171).
If you look carefully at what Telford says there, it’s a truism or a tautology: if the “rate of change is predictable” then you can predict how long ago two species shared a common ancestor. But what if the rate of change is not “predictable”? Apparently, then, all bets are off. And living fossils are a great example of where the rate of evolutionary change was not what was expected — it was much slower. As one paper notes, these “living fossils” often “exhibit slow rates of morphological and molecular change.” So they are apparently one big exception to the rule of predictable rates of evolutionary change.
In reality, as Telford tells us on p. 176: “Genes on different branches of the tree of life have evolved at different rates.” This variation creates a serious challenge for scientists: which lineages or specific genes should serve as the basis for calibrating the clock?
Telford acknowledges this issue directly, noting that “molecular clock estimates sometimes give dates for past events that seem to fly in the face of the fossil evidence” (p. 176). My colleagues, citing other literature, have talked about this previously here. For “living fossils” presumably it ticks very slowly. This issue undermines the molecular clock estimates because it means that scientists must arbitrarily select from which data to calibrate.
- Prediction 3: More similar common characters mean closer relatedness.
A core prediction of evolutionary theory, and a foundational assumption for constructing the tree of life, is that similarity in phenotypic traits (and especially in underlying molecular characters like genes and proteins) reflects common ancestry. As one paper states, “molecular systematics is (largely) based on the assumption … that degree of overall similarity reflects degree of relatedness.” Closely related species should share more similarities due to inheritance from a recent common ancestor, while distant ones diverge more.
However, as any evolutionary biologist will tell you, this has many exceptions! And the exceptions have cleverly also been framed as a prediction of evolutionary theory with the term “convergent evolution.” But convergent evolution was and is not predicted on the hypothesis of common descent. It shows that similarity does not necessarily mean closer relatedness. I’d argue convergent evolution is in fact a strong signal that something is wrong with common descent.
Just how common are these exceptions though? As Telford notes, “convergent evolution of new characters and loss of characters — are very common across the tree of life, typically scattered at random across the branches of the tree” (p. 148). Convergent evolution — where traits arise independently — poses an obvious challenge to the general assumption of evolutionary systematics that similarity results from a common ancestor. And if convergent evolution is “very common” then, though Telford doesn’t acknowledge it, the general premise behind the theory of common descent is in peril.
Swifts and Swallows
Telford gives a good concrete example: swifts and swallows. “Swifts and swallows look and behave remarkably similarly, and they resemble each other in other ways too” (p. 49). He continues: “I have to confess that the almost identical forms of the bodies and behavior of the swifts and the swallows meant that until recently I had assumed they were closely related on the tree of life” (p. 50). Telford notes that he’s in good company because Linnaeus also grouped them in the same genus. Yet, swifts are near hummingbirds and nightjars on the tree of life, while swallows belong to the songbirds, closely related to crows, tits, and wrens (p. 51). To explain this the evolutionary biologist must then accept that their shared aerodynamic adaptation must have evolved twice independently, through convergent evolution.
Here are more examples of exceptions to this key prediction:
- Wings for flight evolved completely independently in insects and birds (p. 140).
- Long, legless bodies have arisen repeatedly in reptiles and amphibians (p. 51).
- Streamlined aquatic bodies evolved separately in dolphins, manatees, and extinct ichthyosaurs (p. 51).
The Data Pushing Back
While physical features already provide countless exceptions, molecular data, where evolutionary change is supposed to be most traceable, reveals an even greater mess.
In Darwin’s time, relying mainly on visible traits made a pattern of common descent seem more straightforward, with details left vague. But once we gained detailed genetic sequences, the clean, unified tree largely disintegrated. Referencing these issues, Telford states on p. 133, “This raises the scary question of whether there even is a single tree of life to discover.” And on p. 258, “Knowing that genes occasionally jump from one branch to another tells us that we cannot always rely on genes to tell us about the relationships between species.” I couldn’t agree more. Or when talking about one of the classic icons of evolution, Darwin’s finches, Telford again states: “But for the Galápagos finches, even entire genomes’ worth of data give inconsistent and confusing answers regarding the relationship between species” (p. 139). To my fellow scientists, I want to emphasize: This is the data pushing back. Are we ready to listen? Or will we keep patching up holes in the theory?
Today, researchers must cherry-pick genetic data to recover the expected tree. See my previous series here and here. When all available genetic data are included, the level of congruence across genes is surprisingly low. So low in fact that scientists aren’t even using all the data to build trees. They almost always use a selected subset which most nicely provides their expected output. Over the last 20+ years, a long list of auxiliary explanations has been introduced to account for the mismatches: horizontal gene transfer, incomplete lineage sorting, convergent evolution at the molecular level, and more. The result is that common descent has become a highly flexible framework — one that can “explain” almost any pattern by adding new ad hoc mechanisms. When a theory explains everything, it risks explaining nothing. For me, the ever-growing collection of rescue hypotheses points to a simpler conclusion: the core idea that all organisms descend from a single common ancestor is incorrect.