Complex III is a molecular machine that plays a critical role in the biochemical generation of ATP by pumping protons across the mitochondrial membrane. Observing the extreme precision and sophistication of Complex III and other molecular machines raises a fundamental question: is it reasonable to attribute their origin to random mutation and natural selection? For natural selection to construct such a system, there would need to be a series of incremental, functional intermediates, each providing a selective advantage. Does such a plausible stepwise pathway exist? Let’s consider that question and others related to the origin of Complex III.
Key Players in the Cell’s Power Plant
Traditionally, five major enzyme complexes are described by what biochemists call the mitochondrial respiratory chain. Engineers would call the mitochondrial respiratory chain a “power plant” where current flows through a circuit to build up voltage. That voltage then spins a nanoscale generator to manufacture the energy currency of life: ATP.
The first protein in the mitochondrial respiratory chain, Complex I, is a jaw-dropping molecular machine I’ve written about before, here, and is a key player in creating the mitochondria’s electrical field. The second enzyme complex, Complex II, acts by accepting electrons from a different molecule, succinate, passing these through a molecule wire which includes the specialized molecule, flavin adenine dinucleotide (FAD or Vitamin B2), and three subsequent iron sulfur clusters (stepping stones for electrons), before finally handing off these electrons to ubiquinone (CoQ). The third enzyme, Complex III, is our subject today. It grabs electrons from the electron shuttle and passes them along, helping to build the proton gradient. Cytochrome c oxidase (Complex IV) receives electrons from mobile cytochrome c proteins, which shuttle one electron at a time from Complex III. It then transfers these electrons to oxygen, the final electron acceptor in oxidative phosphorylation. The fifth is ATP synthase itself — the turbine that turns the proton voltage across the membrane into ATP.
Complex III Basics
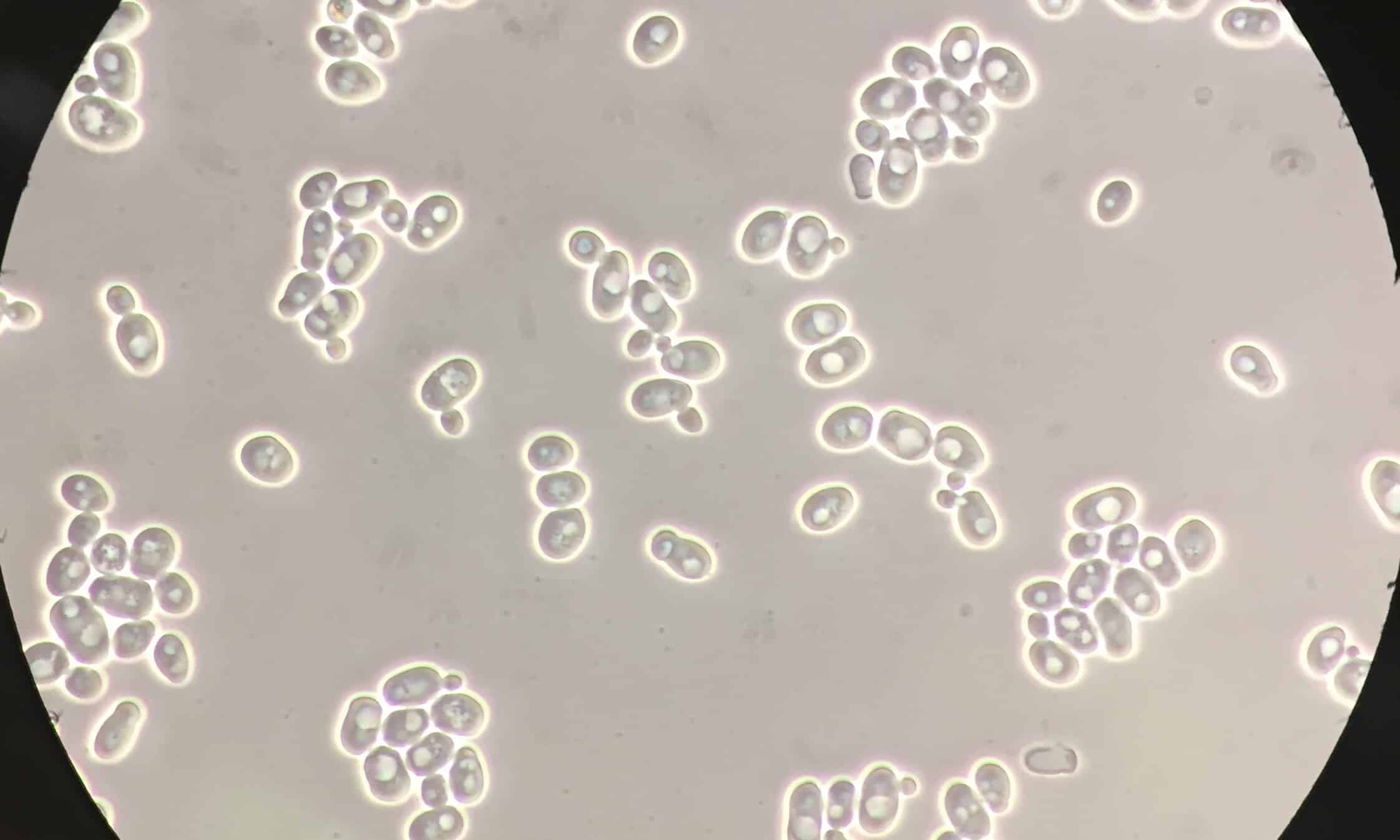
To say that Complex III (also called bc1) is an engineering marvel is an understatement. Complex III is composed of 11 distinct subunits, one of which is encoded by mitochondrial DNA and the remaining ten by nuclear DNA. These 11 subunits assemble into a monomer, but the functional form of the enzyme is a dimer, consisting of 22 subunits in total (Buckel and Thauer 2018). Below is an illustration of Complex III from Saccharomyces cerevisiae, aka brewer’s yeast.
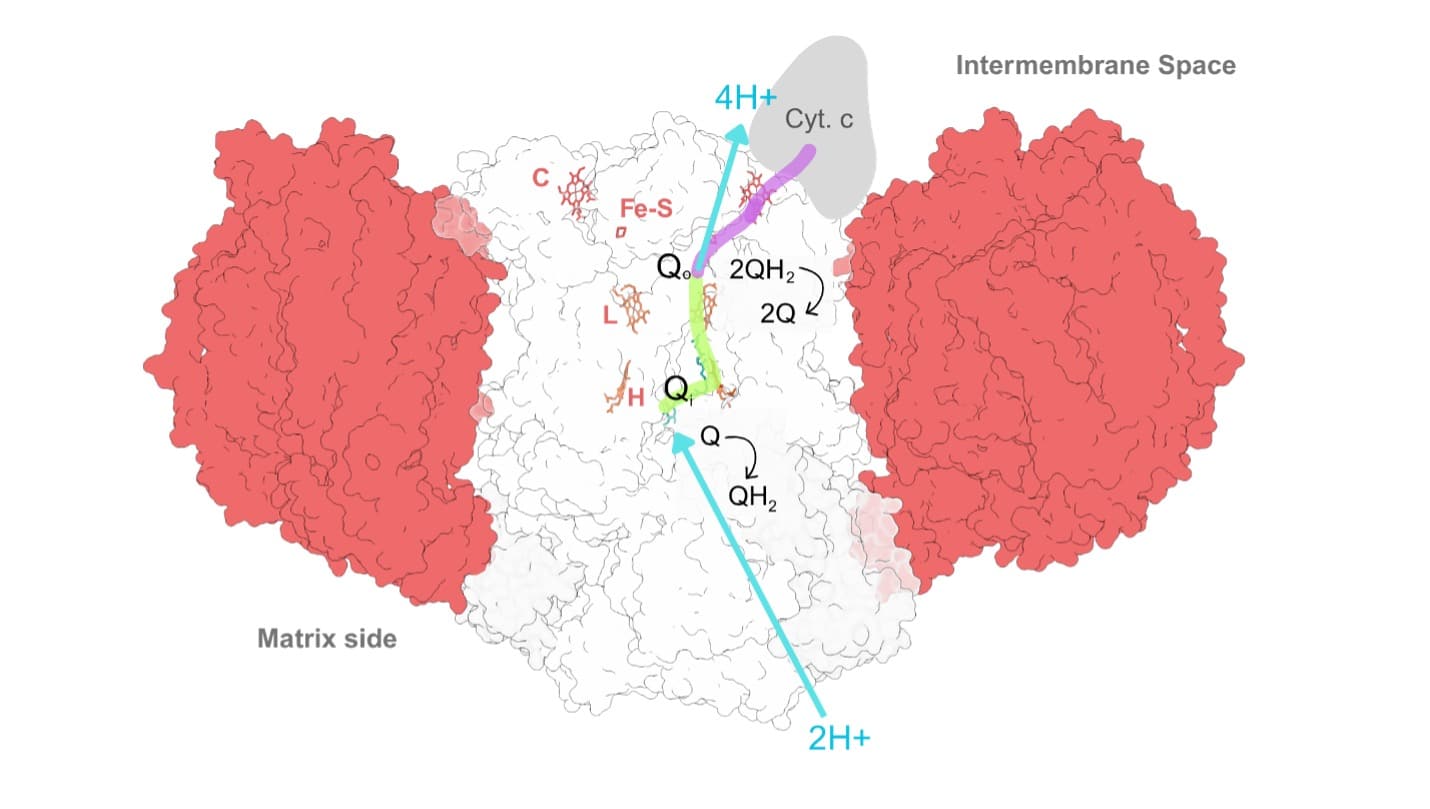
How does it work? Complex III receives two electrons when the mobile electron carrier ubiquinol (QH₂) binds at the Q₀ site. It then sends one electron along a high-energy path (the purple path) to cytochrome c, which carries one electron at a time onward to Complex IV. The second electron follows a lower energy, more “uphill” path and reduces a second molecule of ubiquinone (Q1) bound at the Qᵢ site on the opposite side of the complex (green path). This molecule has the unique and essential ability to hold one electron (a free radical); in this state the molecule is called a semiquinone.
Once the two electrons have been transferred, the two protons originally carried by ubiquinol are released into the intermembrane space. A second molecule of QH₂ then binds at the Q₀ site. Again, one electron takes the high-energy path to cytochrome c, while the second electron travels the low-energy path and further reduces the semiquinone molecule bound at the Qᵢ site to ubiquinol (QH₂). This step draws two protons from the mitochondrial matrix.
In summary, acting like an electron traffic cop, Complex III directs half of the electrons toward cytochrome c (and ultimately ATP production) and the other half toward building a stronger proton gradient across the membrane. Cytochrome c can only accept and carry one electron, so without this bifurcation mechanism one electron would be wasted, leading to heat buildup, in addition to which this free radical could also cause damage to the system. This elegant bifurcation mechanism greatly enhances the overall energy efficiency of the mitochondrial respiratory chain.
Engineering Prowess in the Utilization of Heme
Let me point out some of the engineering prowess here. Complex III relies on three carefully positioned heme molecules: heme b_L, heme b_H, and heme c₁ to accomplish its sophisticated electron routing. Heme is a specialized iron-containing molecule that participates in handling oxygen and electrons. At the center of each heme molecule is an iron atom. This iron cycles between the Fe²⁺ (reduced) and Fe³⁺ (oxidized) states as the electrons move through it. See here for an image of the structure of heme b.
Hurdles for the Origin of Heme
Heme molecules are themselves highly specialized structures, not something that arises for free in biochemistry or on a prebiotic earth. The biosynthesis of heme requires at least eight enzymatic steps, beginning with succinyl-CoA and ending with the final heme molecule. Each step is catalyzed by a dedicated enzyme. Furthermore, the early intermediates of the heme biosynthetic pathway are highly toxic to cells. This inherent toxicity makes stepwise tinkering or gradual assembly a very dangerous process. Selection pressure would strongly favor organisms with a fully functional heme biosynthesis pathway. However, the pathway consists of multiple enzymatic steps, and it is not obvious how natural selection at the organismal level could drive the gradual accumulation of the many mutations required to either repurpose preexisting enzymes into the necessary heme biosynthetic enzymes or to evolve those enzymes entirely de novo.
Electron Movement in Complex III: Reduction Potentials and Quantum Tunneling
Electrons flow “downhill,” which means they flow from more negative reduction potentials to more positive reduction potentials. In Complex III, CoQH2 (reduction potential = +90 mV) gives up its electrons ultimately to a semiquinone and cytochrome c (reduction potential = +250 mV). Reduction potentials are not intrinsic fixed properties of the molecule alone. They depend strongly on the local chemical environment. Complex III’s architecture also plays a key role in making sure the electrons go down physically separate paths, as the Qₒ binding site points the two electrons down different paths.
In Complex III, the high-energy electron travels to the Rieske iron-sulfur (2Fe-2S) cluster (reduction potential = +285 mV), which serves as a stepping stone (Buckel and Thauer 2018) on its pathway toward heme c₁. This is the final stop within the complex before it is transferred to the mobile, soluble cytochrome c. This reaction process is the exergonic (i.e., releases free energy) step that drives the uphill transfer of the second electron. The type of electron movement that happens here is quantum tunneling rather than classical hopping. The key distinction is that classical hopping is a thermally activated process that requires electrons to gain enough energy to hop over an energy barrier, whereas quantum tunneling is a quantum mechanical phenomenon that allows electrons to pass through the energy barrier even when they lack sufficient energy to go over it.
Coenzyme Q (ubiquinone, or CoQ) is a unique redox cofactor that possesses crossed (or inverted) reduction potentials. This property makes the second electron transfer during bifurcation at the Qₒ site of Complex III thermodynamically more accessible. In other words, energetically it is more favorable.
In a typical two-electron redox carrier, removal of the first electron from the fully reduced form produces a semiquinone intermediate whose subsequent oxidation occurs at a more positive reduction potential than the overall two-electron couple. In contrast, for CoQ and similarly for other electron-bifurcating cofactors such as FAD, the situation is reversed. The one-electron semiquinone (Q / SQ) has a more negative reduction potential. As a result, after the first electron is transferred from CoQH2 to the high-potential chain, the remaining semiquinone becomes more willing to give up its electrons, driving the second electron “uphill” along the less favorable low-potential path (to heme b_L).
So in Complex III, the second electron goes from the semiquinone (SQₒ), to heme b_L (reduction potential = −60 mV) (Buckel and Thauer 2018). From heme b_L, the electron then tunnels a distance to heme b_H. Next, it tunnels to a waiting ubiquinone molecule at the Qi site. This initially forms a semiquinone intermediate and ultimately leads to the uptake of protons from the mitochondrial matrix. This pumping of protons contributes to building the proton gradient across the mitochondrial inner membrane. This proton potential is what drives ATP production in Complex V.
Electrochemistry and Quantum Tunneling Requirements for Complex III
If I put on my reverse-engineering hat, this enzyme looks like the work of a designer who deeply understood electrochemistry and quantum tunneling. It appears that the designer of this enzyme knew the ins and outs of electrochemistry and how to exploit quantum tunneling for maximum efficiency. That knowledge is evident in observations such as the precision spacing between the 2Fe-2S cluster and the cytochrome c docking site, what reduction potentials these molecules should have, which amino acids could safely surround the heme molecules without disrupting quantum tunneling, and many other fine details.
Construction of the Rieske 2Fe-2S Cluster
Building the Rieske 2Fe-2S cluster happens in the mitochondria where the atoms come together in the mitochondrial matrix. There is a scaffold protein upon which the sulfur and iron clusters are added. Transfer proteins help to move or convert the cluster. In humans, once the cluster is made, there is a dedicated assembly protein called BCS1L which helps to move the pre-formed cluster into part of Complex III. The assembly protein requires ATP and specifically recognizes a part of Complex III.
Hurdles for the Origin of 2Fe-2S Cluster in Complex III
When a key functional component of a protein is produced outside the protein itself through a separate biosynthetic process and then assembled using a specialized assembly protein, any explanation for the origin of the final protein complex must also account for these preliminary processes. In the case of Complex III, the 2Fe-2S cluster is synthesized externally and must be inserted by a dedicated chaperone protein that specifically interacts with the complex. This means that explaining the origin of Complex III necessarily requires explaining the origin of its 2Fe-2S cluster biosynthesis pathway and the specialized insertion machinery. Accounting for the coordinated evolution of multiple separate proteins and multi-step processes through a blind, unguided mechanism, one lacking any foresight, remains a significant challenge that has yet to be successfully addressed.
Construction of CoQ
Coenzyme Q (ubiquinone) consists of two main structural components that are synthesized through entirely separate biosynthetic pathways: a long hydrophobic polyisoprenoid tail and a quinone (benzoquinone) head group. The hydrophobic tail is constructed via the mevalonate pathway (six steps), which generates isopentenyl pyrophosphate (IPP). These building blocks are then joined together by the PDSS1/PDSS2 complex via nine or ten condensation reactions to form a decaprenyl (C50) chain.
The quinone head group is built from the amino acid tyrosine, which is converted to 4-hydroxybenzoate (4-HB) through a series of reactions (at least three to five steps). Once both precursors are available, the polyisoprenoid tail is attached to the 4-HB ring by the enzyme COQ2. Subsequent modifications to the ring including hydroxylations, methylations, and decarboxylation are carried out by a series of enzymes: COQ3, COQ4, COQ5, COQ6, COQ7, COQ8, COQ9, and COQ10. These proteins work together within the mitochondrial inner membrane to assemble the highly complex CoQ molecule.
Hurdles for Origin of CoQ
Building CoQ is no small task, requiring a minimum of 19 enzymes. Mutations in any of the COQ2 through COQ10 genes, which encode enzymes involved in the final modification steps, can result in severe mitochondrial disease. CoQ is essential for the function of Complex III in the mitochondrial electron transport chain. However, explaining the origin of a functional Complex III also requires explaining the origin of this highly specialized electron-bifurcating cofactor. The intricate biosynthesis and precise biochemical properties of CoQ strongly suggest it is finely tuned for its critical role as a swinging molecule with multiple electron reduction potentials.
The Chemical Brilliance of CoQ
CoQ itself is a masterpiece of organic chemistry. As noted earlier, CoQ10 belongs to a special family of bifurcating cofactors that are capable of stably holding one or two electrons usinga free radical intermediate, which is stabilized through delocalized π-electrons and conjugated double bonds. This structure allows the molecule to contain multiple reduction potentials.
The long isoprenoid tail is precisely the right length to embed the CoQ molecule deeply within the mitochondrial inner membrane. It anchors the quinone head at the optimal depth and orientation, enabling it to diffuse laterally and “swing” just the right distance between the active sites of the respiratory chain complexes.
The quinone head group is exquisitely designed to stabilize a free radical intermediate and manage electrons in a controlled, stepwise manner. It can accept up to two electrons (and two protons) overall, but one at a time.
Coq Picks Up Electrons One at a Time
When the first electron arrives (for example, from the iron-sulfur clusters in Complex I or II), it forms a semiquinone radical anion. The radical is highly stabilized by resonance across the conjugated ring system. A negative charge develops on one of the electronegative oxygen atoms, making it a strong base capable of abstracting a proton from the surrounding environment. The unpaired electron remains delocalized and stabilized within the ring structure, forming the relatively stable semiquinone intermediate.
Coq Hands Electrons Off One at a Time
The second electron then pairs with the radical, fully reducing the molecule to ubiquinol (CoQH₂). This process is reversed at Complex III during the Q-cycle: CoQH₂ donates its electrons one at a time: first to the high energy path, then to the lower energy path which releases protons into the intermembrane space. This stepwise electron handling, combined with precise membrane positioning, allows CoQ to function as a highly efficient, mobile electron carrier.
More Hurdles for the Design of CoQ
Designing the resonance-stabilized quinone ring of CoQ requires a sophisticated understanding of organic chemistry. In addition, the role of electron bifurcation in maximizing energy efficiency must have been appreciated. Synthesizing a molecule with these precise properties through a coordinated pathway involving approximately 19 enzymatic steps strongly suggests the involvement of foresight and purposeful design.
Efficiency in Complex III
Complex III, like many enzymes, makes possible the impossible. The movement of electrons and their charge balancing protons across a distance is a thermodynamically unfavorable reaction, so direct coupling to an exergonic reaction is required to make these processes favorable. How does Complex III do this? As we’ve already discussed, it employs electron bifurcation via the specialized molecule CoQ10.
Let’s talk a little about the efficiency of this electron-bifurcating design. In a comparison of three coupling modes for FAD (see Table X, information taken from (Müller et al. 2018)) electron bifurcation was found to be the most efficient. Both ATP hydrolysis and reverse electron transport require a higher energy cost. Although this was done with FAD, the same principle applies to CoQ10.
| Mode of Coupling | Energy Cost in mV |
| Electron Bifurcation | .23 mol ATP |
| Reverse Electron Transport | 0.58 mol ATP |
| ATP hydrolysis | 1 ATP |
Hurdles for a Darwinian Process Against Thermodynamic Barriers
This type of design is brilliant from an engineering perspective, and it is not the type of design that can be accomplished by blind selection, since two coupled reactions are required to minimize energy loss. In reaction coupling one must be able to understand the total chemical reaction and to conceptualize the complex machinery necessary to carry out the reaction. This requires foresight, something the evolutionary process lacks. How is it proposed that evolution could have added two chemical reactions together, “realized” that the complete chemical reaction would be advantageous, and then start slowly moving toward that? The only way is if each individual part had some selective advantage independently and then somehow they came to work together.
Conclusion
When we observe extreme precision and sophistication in a molecular machine, what does this imply about its origin? Is it reasonable to assume that such a system arose through random mutation and natural selection? For natural selection to build a machine like Complex III, there would need to be a series of incremental, functional improvements at every step of its construction. Yet there appears to be no plausible stepwise pathway, because the primary selective advantage would only emerge once the entire complex, and indeed the full mitochondrial respiratory chain, was already operational.
While it is always possible to invent other scenarios, in the absence of any supporting evidence for those hypothetical intermediates we are left to explain the system as it actually exists.
For Complex III, the picture seems particularly clear. The only process we know of that can design and integrate multicomponent systems, precisely arranging their individual parts and ensuring they function together from the outset, is top-down, goal-directed design by an intelligent agent. Therefore I invite my skeptics to consider honestly: Could a blind, unguided process truly produce a molecular machine of this complexity and elegance?
References
- Buckel, Wolfgang, and Rudolf K. Thauer. 2018. “Flavin-Based Electron Bifurcation, A New Mechanism of Biological Energy Coupling.” Chemical Reviews 118 (7): 3862–3886.
- Müller, Volker, Nilanjan Pal Chowdhury, and Mirko Basen. 2018. “Electron Bifurcation: A Long-Hidden Energy-Coupling Mechanism.” Annual Review of Microbiology 72 (1): 331–353.