A team of scientists from China leveraged AI to design an enzyme that facilitates recycling of polyurethane! This work was published in the journal Science.
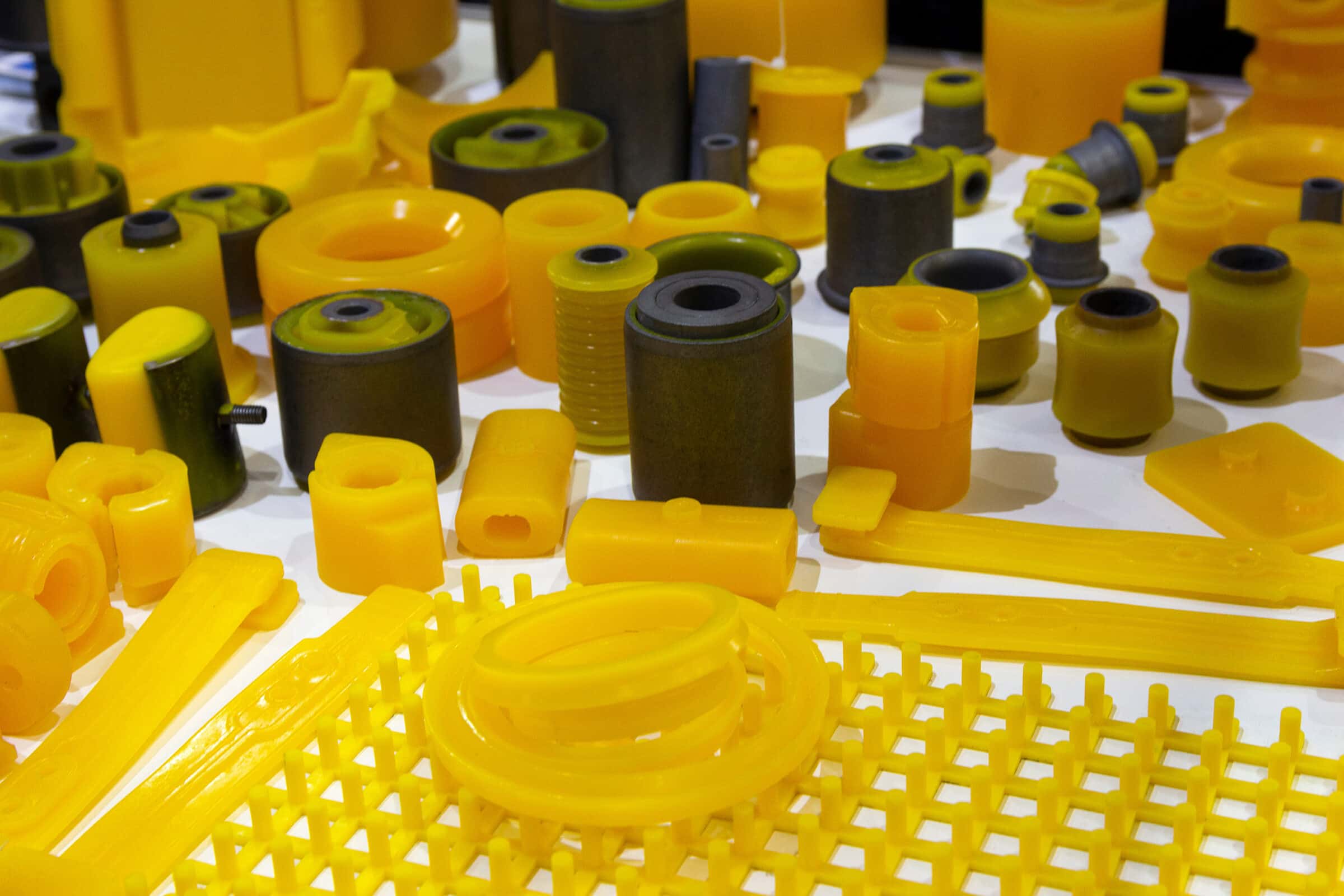
What is polyurethane and where is it used? It’s a type of plastic and it is pretty much everywhere: shoes, clothes, yoga mats, mattresses, couch cushions. Many of these things end up in landfills, where they persist for years, due to its chemical crosslinking.
How Did the Scientists Achieve This?
The researchers developed a process called GRASE (graph neural network–based recommendation of active and stable enzymes) to screen and identify promising urethanases — enzymes that break down urethane bonds in polyurethane. Here’s the step-by-step process:
Step 1: Scientists identified enzymes from nature with “promiscuous” activity against polyurethane. In enzymology, “promiscuous” means these proteins aren’t strictly specialized for one substrate but can loosely bind and partially degrade a range of molecules. These natural enzymes gave the scientists ideas about requirements for an enzyme that could degrade polyurethane.
Step 2: For enzymes lacking experimental structures, the researchers used AlphaFold to predict 3D models. These protein structures were then converted into graph representations — vertices for amino acid types, edges for backbone atomic distances — making them ideal inputs for graph neural networks (GNNs).
Step 3: Next a neural network (how AI works) was used to analyze these graphs and generate novel amino acid sequences that were optimized for a set of preset design features thought to facilitate breakdown of polyurethane.
Step 4: The top candidate sequences were expressed in bacteria (in vivo production) and tested for activity against polyurethane substrates.
The result: Among the hits, an enzyme dubbed AbPURase stood out, showing 32-fold greater activity against one substrate and a 62-fold greater activity against another substrate. With further optimization for industrial conditions, AbPURase achieved near-complete depolymerization of kilogram-scale polyurethane foam in just eight hours — far surpassing the natural enzymes.
Why This Approach Beats Traditional Directed Evolution
Directed evolution leverages mutation and artificial selection. The process asks E. coli populations for a solution, where improvements are identified via selection. It’s successful to an extent but limited by population size, mutation rates/mutation trade-offs, and the ability to design an appropriate selection pressure.
This work flips the script a bit by decoding constraints in existing proteins (fold, catalysis, stability) which enable design of requirements for new functions. No blind search required, and the result? A plastic-eating enzyme in one design cycle.
An Interesting Thought Experiment
Could random mutation plus artificial selection bridge the gap from the closest natural enzyme to this AI-optimized one? Someone needs to run that experiment. As we probe more deeply into the limits of mutation and selection versus foresight-driven engineering, these type of experiments will be key.
This feels like the dawn of design-centric protein engineering.