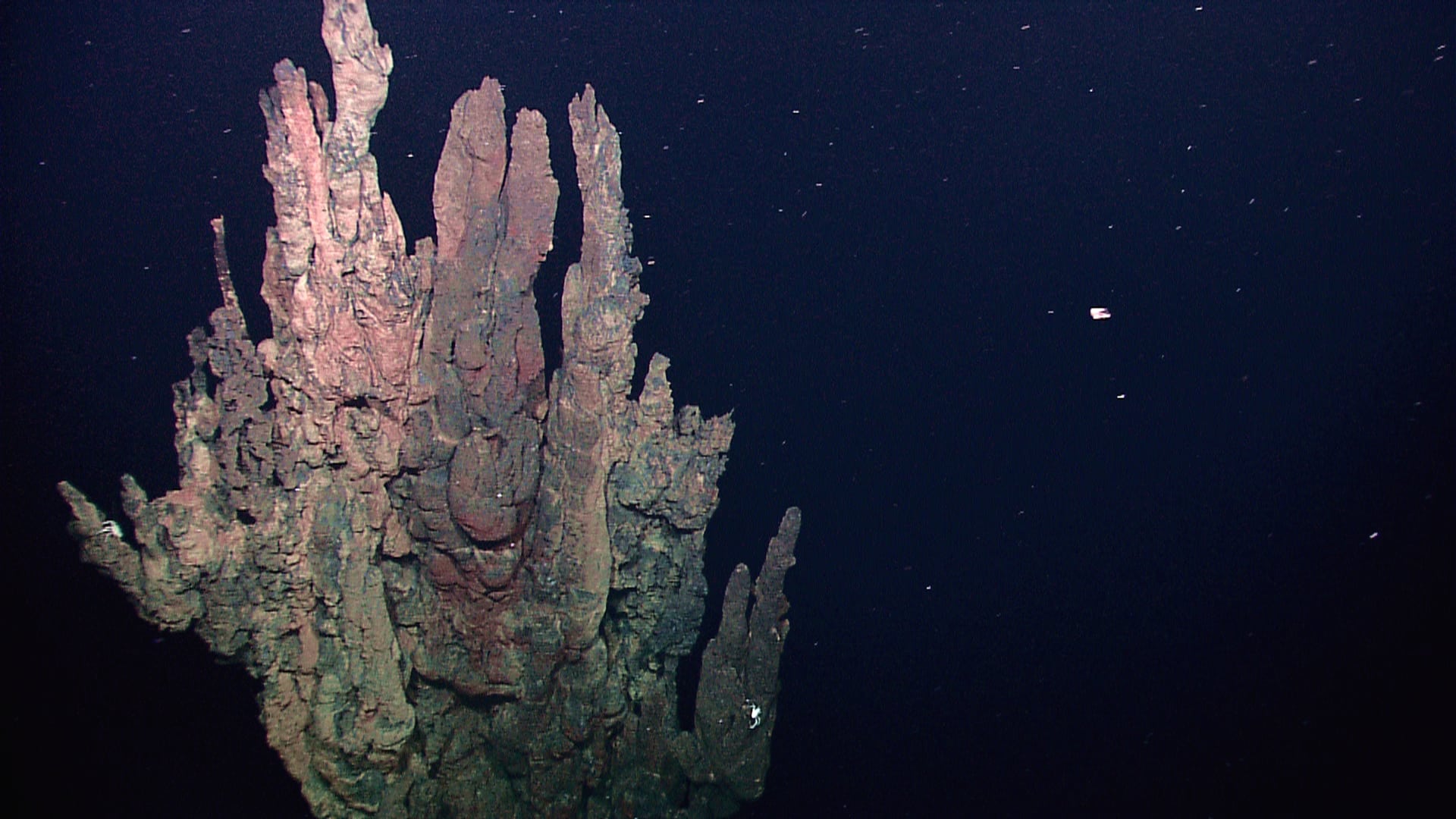
In 2015, a new superphylum of archaea was reported, having been discovered through metagenomic sequences of sediments collected near a hydrothermal vent known as Loki’s Castle in the Arctic Ocean.1 These microorganisms therefore became known as the Asgard archaea. The first characterized lineage was named Lokiarchaeota, and subsequently discovered lineages were named after other Norse deities — Thorarchaeota, Odinarcheota, and Heimdallarchaeota2,3. Phylogenetic analyses of the Asgard genomes, together with the presence of hitherto believed to be “eukaryotic signature proteins” (ESPs) quickly led to these microbes being widely considered to be the closest living prokaryotic relatives of eukaryotes.4,5,6
An Endosymbiotic Relationship?
Nonetheless, there was a problem with this theory. In order for the eukaryotic domain to have arisen from within the Asgard archaea, one of those microbes would need to have engulfed an Alphaproteobacterium and formed an endosymbiotic relationship with it (it is believed that this ultimately gave rise to the mitochondria, which are distinctive of the eukaryotic cell). But the Asgard archaea had been found in anoxic environments — with no or little oxygen. On the other hand, the Alphaproteobacteria are obligate aerobes, requiring oxygen to survive. How, then, could the Asgard archaea and Alphaproteobacteria have met?
A new paper, published in Nature, offers a plausible solution to this enigma.7 This new study determined that,
Heimdallarchaeia form metabolic guilds that are distinct from other Asgardarchaeota. These archaea encode hallmark proteins of an aerobic lifestyle, including electron transport chain complex (IV), haem biosynthesis and reactive oxygen species detoxification. Heimdallarchaeia also encode novel clades of respiratory membrane-bound hydrogenases with additional Complex I-like subunits, which potentially increase proton-motive force generation and ATP synthesis.
While this analysis does resolve one enigma (though a relatively trivial one given our limited sampling), there remains a far greater problem for the postulate that we eukaryotes owe our origins to the Asgard archaea. The modes of cell division of those two types of cells are wildly different.
How Do Archaea Divide?
Cell division in archaea has been much less well studied than in bacteria. Nonetheless, it is known that some archaea (in particular the Euryarchaeota) divide in a manner very much like bacterial division (i.e., FtsZ-based division).8 This mechanism involves the polymerization of FtsZ proteins into filaments and their assembly around the division septum. This ring then recruits further division proteins. Membrane constriction ultimately brings about two daughter cells.
A more common mechanism of cell division among archaea is ESCRT-III division.9,10,11 These proteins exhibit homology to the machinery involved in cytokinesis, membrane scission and endosomal sorting in eukaryotic cells. Proteins called Cdv proteins assemble at the midpoint of the cell. A constriction ring made of ESCRT-III filaments forms. These filaments are disassembled by the ATPase CdvC. Finally, division is completed by membrane scission. Though this archaeal mode of cell division more closely resembles eukaryotic cell division than does bacterial cell division, there are still very major differences. Most importantly, archaea do not possess the elaborate mitotic machinery found in eukaryotes. Indeed, ESCRT proteins in eukaryotes are only involved in the final stage of cytokinesis, not chromosome segregation. Chromosome segregation occurs in a manner resembling the bacterial partitioning system involving ParB (which binds to the parS site near the replication origin) and ParA (which binds nonspecifically across the chromosome, having been recruited by ParB).12,13 ParA possesses ATPase activity. Hydrolysis of ATP creates a dynamic gradient, resulting in the chromosome origins migrating toward opposite poles of the cell.
Archaeal Division Is Wildly Different from Eukaryotic Mitosis
Archaeal chromosome segregation looks nothing like eukaryotic mitosis. Microtubules, though an absolutely essential aspect of mitosis, are entirely absent from the archaeal division process. Archaea also lack the actin-myosin contractile ring found in eukaryotes. The tight regulation found in mitotic cell division is also absent from archaeal division, which has a much simpler control system. Though the ESCRT-III system (found in many archaea, including Asgard, where it forms the main cytokinesis machinery) is homologous to the eukaryotic equivalent, it plays only a minor role in mitotic division, performing the final membrane cut that separates the two daughter cells.
Eukaryogenesis: Still an Enigma
Given the presence of previously believed-to-be eukaryotic signature proteins among the Asgard archaea, this superphylum represents the most probable prokaryotic ancestor of eukaryotes, conditional on the assumption that such an ancestor existed at all. For the hypothesis that eukaryotes emerged from within the Asgard archaea, however, it must be demonstrated that a process plausibly existed for transitioning one cell division system into a fundamentally different apparatus for bringing about division, without passing through any maladaptive intermediate stages. Not only this, but the components themselves, which are known to be critical for successful mitosis, must arise de novo, since the vast majority of mitotic components do not have homologues among prokaryotes, including the Asgard archaea.14 Unless and until a plausible solution is offered to this obstacle, serious doubts remain about whether such a transition even occurred.
Notes
- Spang A, Saw JH, Jørgensen SL, Zaremba-Niedzwiedzka K, Martijn J, Lind AE, van Eijk R, Schleper C, Guy L, Ettema TJG. Complex archaea that bridge the gap between prokaryotes and eukaryotes. Nature. 2015 May 14;521(7551):173-179. doi: 10.1038/nature14447. Epub 2015 May 6. PMID: 25945739; PMCID: PMC4444528.
- Seitz KW, Lazar CS, Hinrichs KU, Teske AP, Baker BJ. Genomic reconstruction of a novel, deeply branched sediment archaeal phylum with pathways for acetogenesis and sulfur reduction. ISME J. 2016 Jul;10(7):1696-705. doi: 10.1038/ismej.2015.233. Epub 2016 Jan 29. PMID: 26824177; PMCID: PMC4918440.
- Zaremba-Niedzwiedzka K, Caceres EF, Saw JH, Bäckström D, Juzokaite L, Vancaester E, Seitz KW, Anantharaman K, Starnawski P, Kjeldsen KU, Stott MB, Nunoura T, Banfield JF, Schramm A, Baker BJ, Spang A, Ettema TJ. Asgard archaea illuminate the origin of eukaryotic cellular complexity. Nature. 2017 Jan 19;541(7637):353-358. doi: 10.1038/nature21031. Epub 2017 Jan 11. PMID: 28077874.
- Köstlbacher, S., van Hooff, J.J.E., Panagiotou, K. et al. Prediction of eukaryotic cellular complexity in Asgard archaea using structural modelling. Nat Microbiol 11, 747–758 (2026). https://doi.org/10.1038/s41564-026-02273-y
- Tobiasson V, Luo J, Wolf YI, Koonin EV. Dominant contribution of Asgard archaea to eukaryogenesis. bioRxiv [Preprint]. 2025 Jul 14:2024.10.14.618318. doi: 10.1101/2024.10.14.618318. Update in: Nature. 2026 Feb;650(8100):141-149. doi: 10.1038/s41586-025-09960-6. PMID: 40791505; PMCID: PMC12338589.
- Albers S, Ashmore J, Pollard T, Spang A, Zhou J. Origin of eukaryotes: What can be learned from the first successfully isolated Asgard archaeon. Fac Rev. 2022 Jan 27;11:3. doi: 10.12703/r-01-000005. PMID: 35174363; PMCID: PMC8815363.
- Appler KE, Lingford JP, Gong X, Panagiotou K, Leão P, Langwig MV, Greening C, Ettema TJG, De Anda V, Baker BJ. Oxygen metabolism in descendants of the archaeal-eukaryotic ancestor. Nature. 2026 Feb 18. doi: 10.1038/s41586-026-10128-z. Epub ahead of print. PMID: 41708851.
- Ithurbide S, Gribaldo S, Albers SV, Pende N. Spotlight on FtsZ-based cell division in Archaea. Trends Microbiol. 2022 Jul;30(7):665-678. doi: 10.1016/j.tim.2022.01.005. Epub 2022 Mar 1. PMID: 35246355.
- Liu J, Lelek M, Yang Y, Salles A, Zimmer C, Shen Y, Krupovic M. A relay race of ESCRT-III paralogs drives cell division in a hyperthermophilic archaeon. mBio. 2025 Feb 5;16(2):e0099124. doi: 10.1128/mbio.00991-24. Epub 2024 Dec 19. PMID: 39699168; PMCID: PMC11796394.
- Caspi Y, Dekker C. Dividing the Archaeal Way: The Ancient Cdv Cell-Division Machinery. Front Microbiol. 2018 Mar 2;9:174. doi: 10.3389/fmicb.2018.00174. PMID: 29551994; PMCID: PMC5840170.
- Samson RY, Dobro MJ, Jensen GJ, Bell SD. The Structure, Function and Roles of the Archaeal ESCRT Apparatus. Subcell Biochem. 2017;84:357-377. doi: 10.1007/978-3-319-53047-5_12. PMID: 28500532.
- Kalliomaa-Sanford AK, Rodriguez-Castañeda FA, McLeod BN, Latorre-Roselló V, Smith JH, Reimann J, Albers SV, Barillà D. Chromosome segregation in Archaea mediated by a hybrid DNA partition machine. Proc Natl Acad Sci U S A. 2012 Mar 6;109(10):3754-9. doi: 10.1073/pnas.1113384109. Epub 2012 Feb 21. PMID: 22355141; PMCID: PMC3309785.
- Yen CY, Lin MG, Chen BW, Ng IW, Read N, Kabli AF, Wu CT, Shen YY, Chen CH, Barillà D, Sun YJ, Hsiao CD. Chromosome segregation in Archaea: SegA- and SegB-DNA complex structures provide insights into segrosome assembly. Nucleic Acids Res. 2021 Dec 16;49(22):13150-13164. doi: 10.1093/nar/gkab1155. PMID: 34850144; PMCID: PMC8682754.
- McLatchie J (2024) Phylogenetic Challenges to the Evolutionary Origin of the Eukaryotic Cell Cycle. BIO-Complexity 2024 (4):1–19 doi:10.5048/BIO-C.2024.4